This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Ovarian Cancer and RAD51D
Ovarian cancer is the deadliest of the gynecological cancers, with an estimated 14,000 women expected to die of the disease in 2019 alone [1]. While the cause of ovarian cancer remains unknown, there are a number of known genetic and environmental risk factors that influence an individual's chance of developing ovarian cancer. For example, some environmental factors, such as having a child before 26, or taking oral contraceptive have been shown to lower an individual's risk [2]. However, certain factors such as obesity, or inheriting genetic risk factors, have been shown to increase an individual's risk [2]. Given the fact that early symptoms are rarely manifested in ovarian cancer, and those that do appear aren't immediately distinguishable from other conditions, catching the disease in its early stages can be difficult [2]. However, in order to give a patient an adequate chance to effectively treat ovarian cancer, it is essential to obtain a diagnosis as soon as possible. Yet, with advances in multi-gene panel testing, women with a family history associated with breast and ovarian cancer can be screened for mutations for a number of genetic risk factors, and by doing so can understand their individual risks for ovarian cancer, and obtain a plan for early intervention.
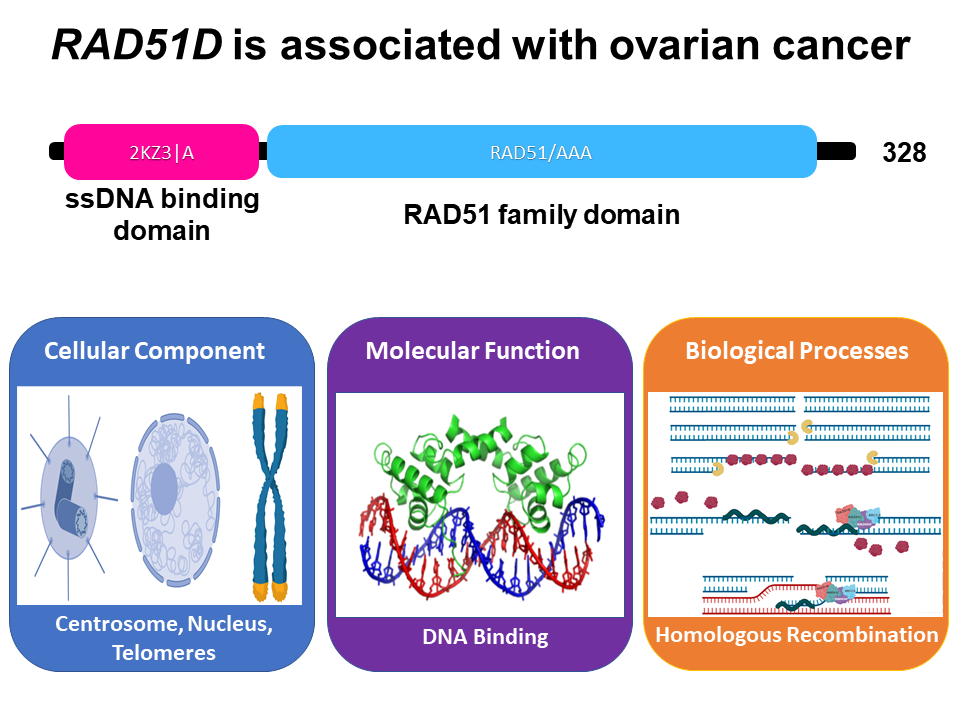
While mutations in the RAD51D gene are rare, they are associated with an increased risk of developing ovarian cancer. A visual depiction of the RAD51D protein can be seen below (Figure 1). There are two protein domains present in the RAD51D protein, both of which are associated with DNA binding. At the N terminus has been shown to preferentially bind single-stranded DNA, allowing the protein to sample for damaged sites of DNA. Additionally, the RAD51 family domain at the C-terminus is present in all RAD51 family proteins, which hydrolyzes ATP in order to "walk" along DNA molecules. Given its molecular function in DNA binding, RAD51D predominately resides in the Nucleus of the cell, and is mainly associated with regulating homologous recombination, both in the context of meiosis and DNA repair. Interestingly, the RAD51 family domain is conserved across all eukaryotes; however, the same is not true for the N-terminus single-stranded DNA binding domain. Instead, the ssDNA binding domain is conserved only in vertebrate model organisms, suggesting that perhaps the ssDNA domain evolved to combat increased instances of DNA damage being induced in the ovaries of vertebrate species.
While mutations in the RAD51D gene are rare, they are associated with an increased risk of developing ovarian cancer. A visual depiction of the RAD51D protein can be seen below (Figure 1). There are two protein domains present in the RAD51D protein, both of which are associated with DNA binding. At the N terminus has been shown to preferentially bind single-stranded DNA, allowing the protein to sample for damaged sites of DNA. Additionally, the RAD51 family domain at the C-terminus is present in all RAD51 family proteins, which hydrolyzes ATP in order to "walk" along DNA molecules. Given its molecular function in DNA binding, RAD51D predominately resides in the Nucleus of the cell, and is mainly associated with regulating homologous recombination, both in the context of meiosis and DNA repair. Interestingly, the RAD51 family domain is conserved across all eukaryotes; however, the same is not true for the N-terminus single-stranded DNA binding domain. Instead, the ssDNA binding domain is conserved only in vertebrate model organisms, suggesting that perhaps the ssDNA domain evolved to combat increased instances of DNA damage being induced in the ovaries of vertebrate species.
Figure 1: A visual depiction of the DNA binding domains in the human RAD51D protein.
Mutations in RAD51D affect DNA Repair
|
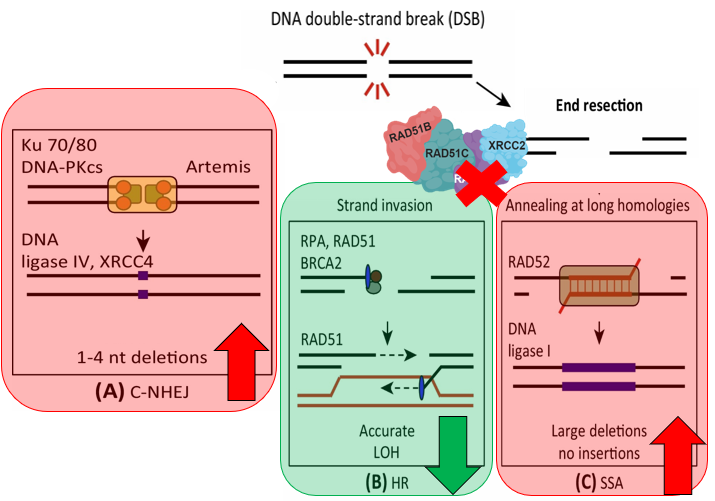
DNA is constantly undergoing damage in mammalian cells, and for the most part, is accurately repaired within the cell. One form of DNA damage, known as a double strand break can be induced due to high-energy radiation, or through stress associated with DNA replication. If left unrepaired, double-strand breaks are lethal for the cell, thus explaining why cells have evolved multiple pathways for repairing these DNA lesions. In fact, RAD51D plays an important role in regulating the early steps of one such DNA repair pathway, known as homology directed repair, which accurately repairs damaged DNA through the use of a homologous template sequence. Therefore, loss of function mutations in the RAD51D protein lead to the inability for the cell to catalyze homology directed repair, causing the recruitment of the deletion-prone DNA repair pathways shown in Figure 2 [3]. Yet, recent research has shown that RAD51D mutations not only affect the use of these alternative pathways, but the deletions incurred by them are significantly larger than in normal wild-type cells, suggesting that RAD51D protects the genome against large-scale deletions [3]. |
Figure 2: Adapted from https://doi.org/10.1016/j.tcb.2015.07.009. Diagram showing the repair pathways utilized in RAD51D knockouts to repair double-stranded breaks.
|
Gap in Knowledge
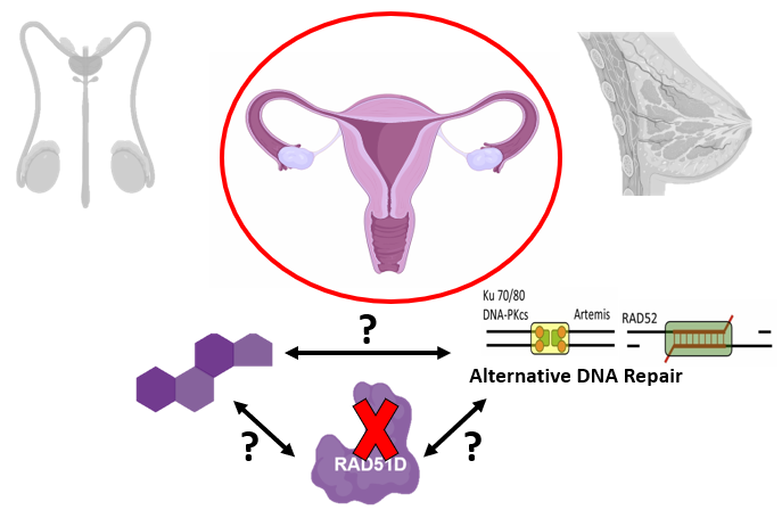
Interestingly, while some studies have found possible associations between RAD51D and breast cancer, recent meta-analyses have failed to reach similar conclusions, showing RAD51D is only a high risk allele for developing ovarian cancer [4,5]. Despite these findings, it is unknown why RAD51D is a tissue-specific risk allele. My primary goal for this research project is to better understand how the RAD51D protein regulates DNA repair in the ovarian microenvironment. My long-term goal is to understand whether or not RAD51D is in fact a tissue-specific risk allele exclusive to ovarian cancer. My hypothesis is that RAD51D has evolved certain features in vertebrate species that are protective against deletions associated with proliferative signals that are present in the ovarian microenvironment.
Figure 3: Diagram illustrating the primary goal of this research endeavor, seeking to better understand RAD51D's role as a tissue-specific risk allele
Model Organisms
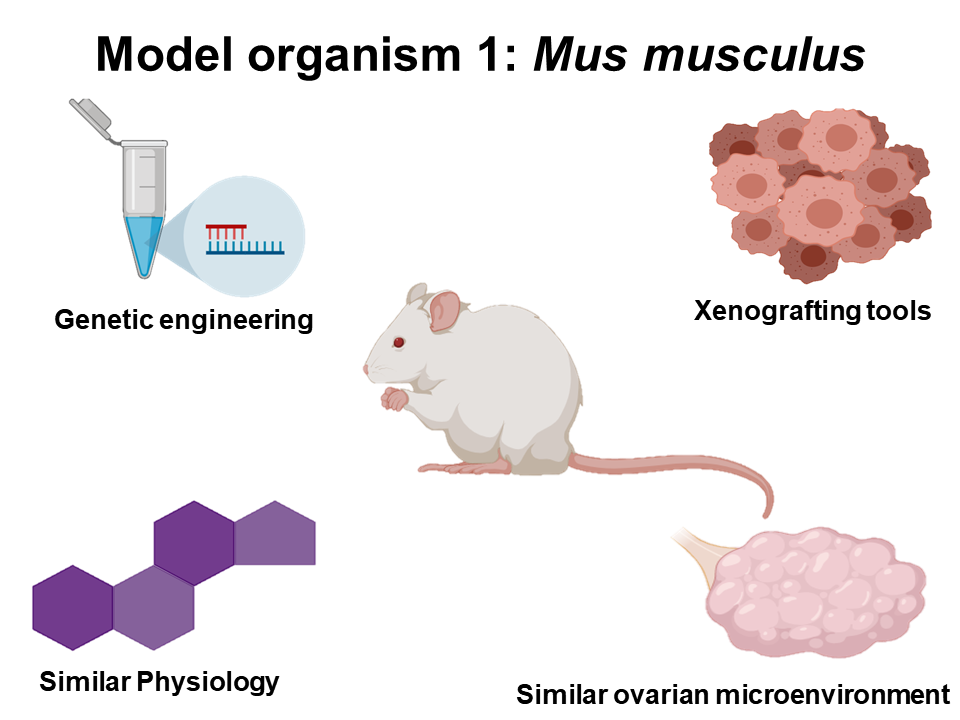
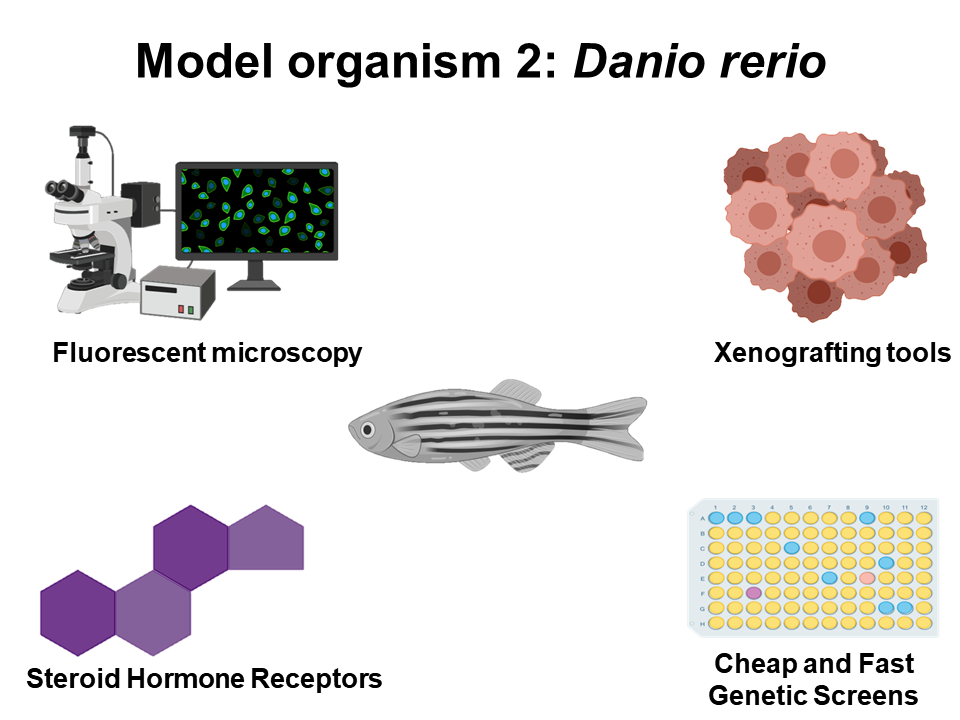
In order to accurately model human ovarian cancer and DNA repair processes, both Mus musculus and Danio rerio will be used for this research project. Ultimately, both of these organisms have been extremely successful models for studying ovarian cancer, and since their ovarian microenvironment is similar to that of human beings, both mice and zebrafish are excellent candidates for this work. Additionally, mice have a plethora of genetic engineering tools and protocols for introducing tissue-specific and developmental-specific mutations. Furthermore both organisms can be used to xenograft human tumor cells, giving researcher the opportunity to study real human tumors in their natural environments. Finally, zebrafish models provide an excellent tool for cheap and fast genetic screens, that when coupled to fluorescent reporters will be important for studying DNA repair.
Figure 4: Diagrams illustrating why the organisms chosen for this project are appropriate for modeling ovarian cancer.
Aim 1: Identify conserved amino acids among vertebrate species
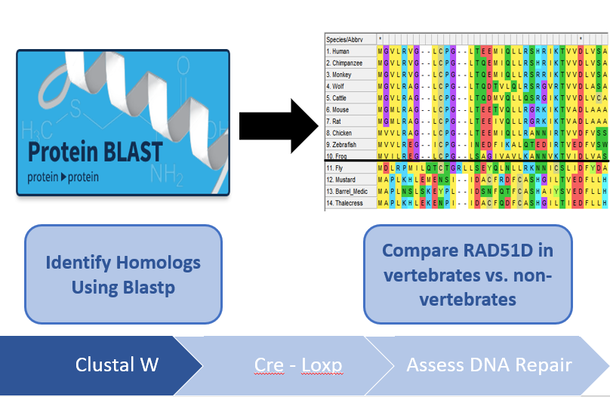
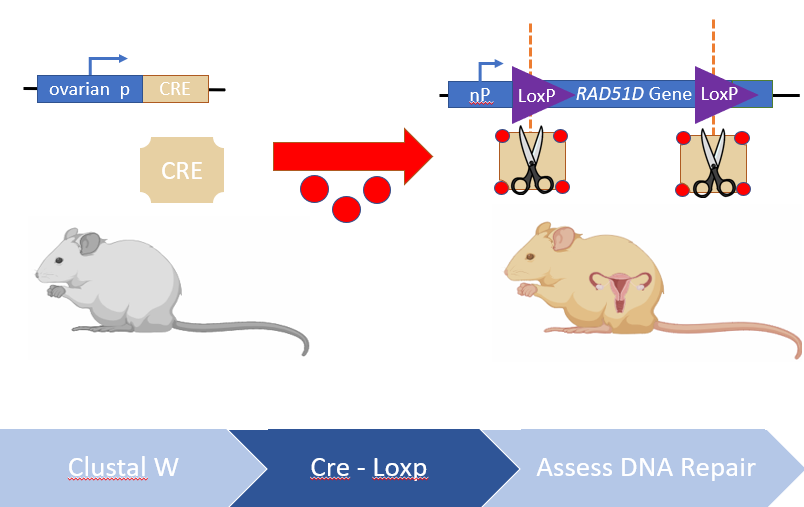
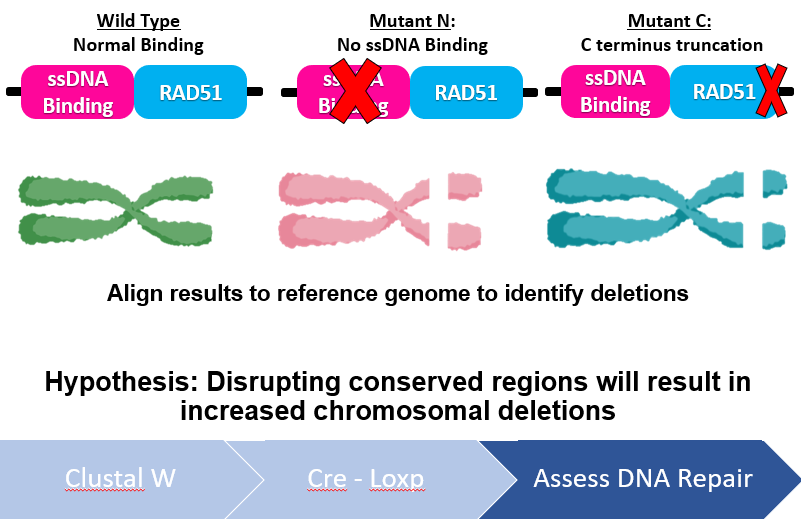
I will use NCBI's BLASTp and ENSEMBL to identify RAD51D protein homologs in different model organisms. By obtaining the protein FASTA sequence of these protein homologs, they can be aligned using MEGA-X's ClustalW algorithm and conserved amino acids among vertebrates vs non-vertebrates can be compared. After analyzing the protein alignments, there were two regions identified in the alignments that were exclusive to vertebrate ovaries, one of which being the N-terminus ssDNA binding protein, and another region towards the end of the C-terminus. Since RAD51D mutations are embryonic lethal, Cre-LoxP constructs will be used to truncate both of the regions identified in the ClustalW alignments in ovarian tissues post-embryonic development. Finally, any tumors that are generated from these transgenic mice or wild type mice will analyzed using next-gen sequencing, to determine if the loss of these regions contribute to large deletions within the ovarian cells. I hypothesize that disrupting conserved regions of the RAD51D protein among vertebrate species will lead to increased chromosomal deletions within ovarian tissues.
Figure 5: Diagram demonstrating the workflow of Aim 1. Conserved regions will be identified using bioinformatic tools, truncated using Cre-LoxP and any deletions incurred analyzed by Next-Gen sequencing.
Aim 2: Identify hormones that affect ovarian genetic integrity
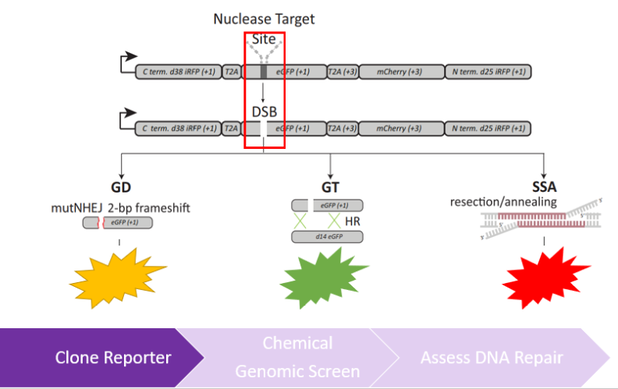
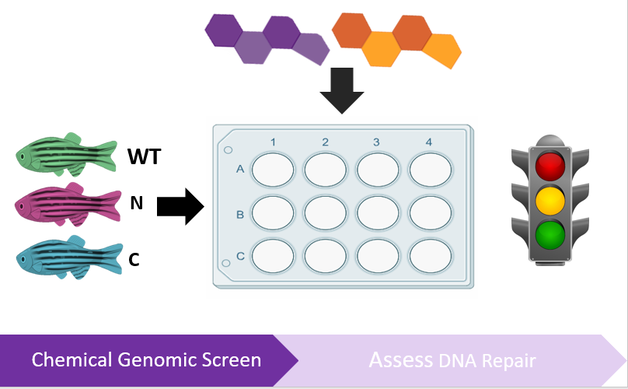
In order to better understand how hormones influence DNA repair in the ovaries, a chemical genetic screen will be used, composed of hormones that important in the ovarian microenvironment, such as estrogen, progesterone, testosterone, and other external growth factors. These hormones will be tested individually and in combination to determine how these hormones regulate different DNA repair pathways. To identify changes in DNA repair, zebrafish will co-transformed with the Traffic Light reporter described in Kuhar et al. 2013 and is shown in Figure 6 [6]. Both wild-type and mutant RAD51D lines of zebrafish will be used to understand how all three of these factors interact in the ovaries. I hypothesize that there will be increased use of the deletion-prone single strand annealing pathway in the RAD51D mutants in response to proliferative signals in the ovaries; however, certain hormones, such as progesterone and testosterone will not have significant effects on regulating DNA repair in the RAD51D mutants.
Figure 6: Diagrams describing the workflow of Aim 2. The left diagram shows the use of the Traffic Light Reporter which can distinguish between different DNA repair pathways used to repair a double-strand break. The right diagram shows the experimental design a genetic screen used to assess DNA repair in the ovaries.
Aim 3: Identify changes in protein-protein binding interactions in RAD51D mutants
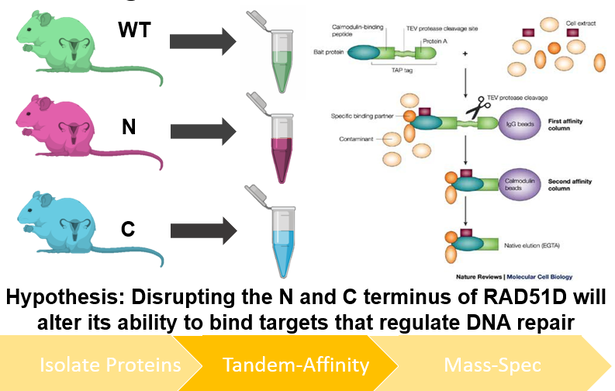
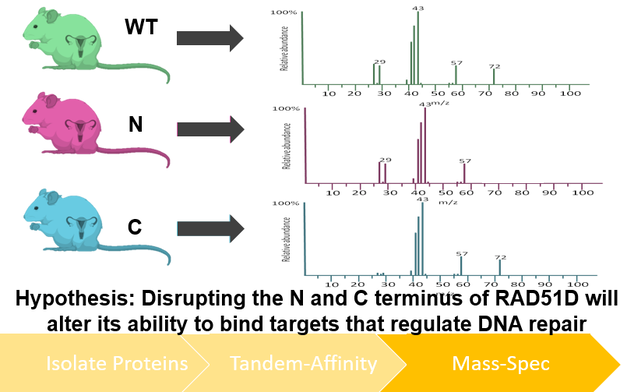
I will use TAP-tagging of proteins isolated from mice ovaries to better understand how the mutants described in Aim 1 regulate DNA repair at the proteomic level. After performing TAP-tag purification of both wildtype and mutant RAD51D isolates, purified protein complexes will be identified using mass spectrometry, and their respective roles in DNA repair characterized using the gene ontology database Panther. I hypothesize that disrupting either the N or C terminus of RAD51D will alter its ability to bind targets that regulate DNA repair.
Figure 7: Diagram depicting the workflow of Aim 3. Proteins will be isolated from both wildtype and mutant RAD51D mice from ovarian tissue and purifyied using TAP tagging (left). Protein interactions within the TAP-tagging isolation process will be identified using mass spectrometry (right).
Future Directions
|
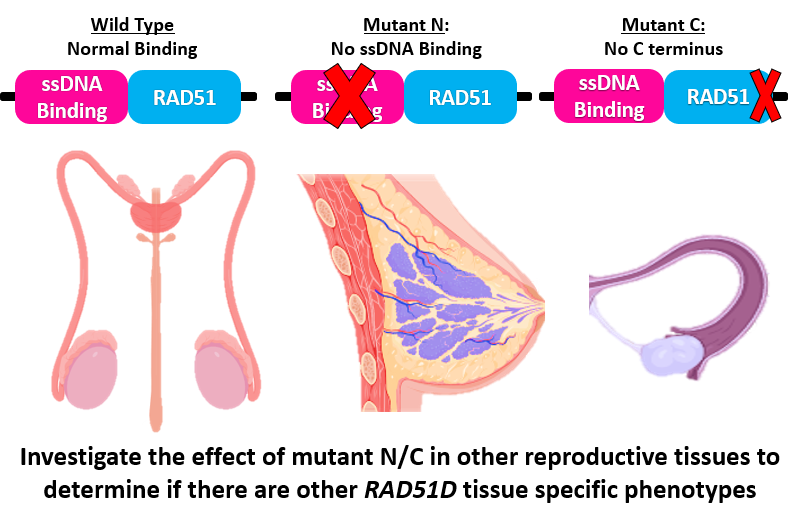
Figure 8: Diagram depicting various tissues that will be targeted during future experiments.
|
The primary goal of this work is to better understand how the RAD51D protein regulates DNA repair in the ovarian microenvironment. However, more research will need to be conducted for the long-term goal of understanding why RAD51D is a tissue specific risk allele. To further this work, the experiments outlined in this proposal will be repeated in other reproductive tissues such as the breast or testes. By doing so, results from different tissues will be compared and contrasted from those found in the ovaries, providing a better understanding of how RAD51D regulates DNA repair in different tissues.
Furthermore, RAD51D may contribute to the development of ovarian cancer external form ovarian tissue. Given the fact that many cases of ovarian cancer actually originate from the Fallopian tubes, this experiment should be repeated within Fallopian tissues to provide a more complete picture of how the loss of the RAD51D protein contributes to the development of ovarian cancer. Ultimately, the results of this experiment, as well as the ones proposed for the future, will allow for both male and female carriers of RAD51D mutations, to better understand their individual risks for developing cancer. |
Rough Draft_1 : https://drive.google.com/drive/folders/1oiQ3FFRLdVEVq7fh4Bli9Hcbso6H2dF8?usp=sharing (More recent below)
| final_talk_jl_draft_3.pptx | |
| File Size: | 4456 kb |
| File Type: | pptx |
| lombardino_jonathan_5_3_2019_final_talk.pdf | |
| File Size: | 3556 kb |
| File Type: | |
References:
1. American Cancer Society. Key Statistics for Ovarian Cancer (2018, April 11). Retrieved from https://www.cancer.org/cancer/ovarian-cancer/about/key-statistics.html
2. Lisa Fayed. Very Well Health. Symptoms of Ovarian Cancer. Retrieved from https://www.verywellhealth.com/ovarian-cancer-causes-risk-factors-2509671
3. Reh, W. A., Nairn, R. S., Lowery, M. P., & Vasquez, K. M. (2016). The homologous recombination protein RAD51D protects the genome from large deletions. Nucleic acids research, 45(4), 1835–1847. doi:10.1093/nar/gkw1204
4. Malwina, S., Katarzyna, K., Jasinska, A., Piotr, K. (2019, in press). Large-scale meta-analysis of mutations identified in panels of breast/ovarian cancer-related genes — Providing evidence of cancer predisposition genes. Gynecologic Oncology. doi: 10.1016/j.ygyno.2019.01.027
5. Kurian, A., Hughes, E., Handorf, E. A., Gutin, A., Allen, B., Hartman, R., Hall, M. J. (2017). Breast and Ovarian Cancer Penetrance Estimates Derived From Germline Multiple-Gene Sequencing Results in Women. JCO Precision Oncology, [1], 1-12. doi: 10.1200/PO.16.00066
6. Kuhar, R., Gwiazda, K. S., Humbert, O., Mandt, T., Pangallo, J., Brault, M., … Certo, M. T. (2013). Novel fluorescent genome editing reporters for monitoring DNA repair pathway utilization at endonuclease-induced breaks. Nucleic acids research, 42(1), e4. doi:10.1093/nar/gkt872
2. Lisa Fayed. Very Well Health. Symptoms of Ovarian Cancer. Retrieved from https://www.verywellhealth.com/ovarian-cancer-causes-risk-factors-2509671
3. Reh, W. A., Nairn, R. S., Lowery, M. P., & Vasquez, K. M. (2016). The homologous recombination protein RAD51D protects the genome from large deletions. Nucleic acids research, 45(4), 1835–1847. doi:10.1093/nar/gkw1204
4. Malwina, S., Katarzyna, K., Jasinska, A., Piotr, K. (2019, in press). Large-scale meta-analysis of mutations identified in panels of breast/ovarian cancer-related genes — Providing evidence of cancer predisposition genes. Gynecologic Oncology. doi: 10.1016/j.ygyno.2019.01.027
5. Kurian, A., Hughes, E., Handorf, E. A., Gutin, A., Allen, B., Hartman, R., Hall, M. J. (2017). Breast and Ovarian Cancer Penetrance Estimates Derived From Germline Multiple-Gene Sequencing Results in Women. JCO Precision Oncology, [1], 1-12. doi: 10.1200/PO.16.00066
6. Kuhar, R., Gwiazda, K. S., Humbert, O., Mandt, T., Pangallo, J., Brault, M., … Certo, M. T. (2013). Novel fluorescent genome editing reporters for monitoring DNA repair pathway utilization at endonuclease-induced breaks. Nucleic acids research, 42(1), e4. doi:10.1093/nar/gkt872
References - Images:
Figure 1: https://biorender.com/
Figure 2: https://doi.org/10.1016/j.tcb.2015.07.009
Figure 3: https://biorender.com/, https://doi.org/10.1016/j.tcb.2015.07.009
Figure 4: https://biorender.com/
Figure 5: https://blast.ncbi.nlm.nih.gov/Blast.cgi , https://www.megasoftware.net/
Figure 6: https://i.pinimg.com/originals/28/ff/6f/28ff6f46d418860eacb64f4d62a86a96.jpg, https://biorender.com/,
https://www.researchgate.net/publication/257754595_Novel_fluorescent_genome_editing_reporters_for_monitoring_DNA_repair_pathway_utilization_at_endonuclease-induced_breaks
Figure 7: https://www.researchgate.net/publication/10964536_Opinion_Is_proteomics_heading_in_the_wrong_direction/figures, https://biorender.com/
Figure 8: https://biorender.com/
Figure 2: https://doi.org/10.1016/j.tcb.2015.07.009
Figure 3: https://biorender.com/, https://doi.org/10.1016/j.tcb.2015.07.009
Figure 4: https://biorender.com/
Figure 5: https://blast.ncbi.nlm.nih.gov/Blast.cgi , https://www.megasoftware.net/
Figure 6: https://i.pinimg.com/originals/28/ff/6f/28ff6f46d418860eacb64f4d62a86a96.jpg, https://biorender.com/,
https://www.researchgate.net/publication/257754595_Novel_fluorescent_genome_editing_reporters_for_monitoring_DNA_repair_pathway_utilization_at_endonuclease-induced_breaks
Figure 7: https://www.researchgate.net/publication/10964536_Opinion_Is_proteomics_heading_in_the_wrong_direction/figures, https://biorender.com/
Figure 8: https://biorender.com/