This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Introduction
What is Gene Ontology?
Imagine finding a particular gene of interest that you would like to learn more about. Some common questions that you would want to answer would likely be what processes is my gene associated with, where in the cell it is located, and what are some of its molecular roles within the cell. One option could be to scour through a long list of publications to find critical information, but as one can imagine, can be a daunting task. Luckily, researchers have access to a useful and organized way of communicating a gene's important information called Gene Ontology (GO). Essentially GO is a collection of defined terms that represent a gene of interest's characteristics or properties. To communicate important information about a gene, GO is broken down into three different aspects: [1]
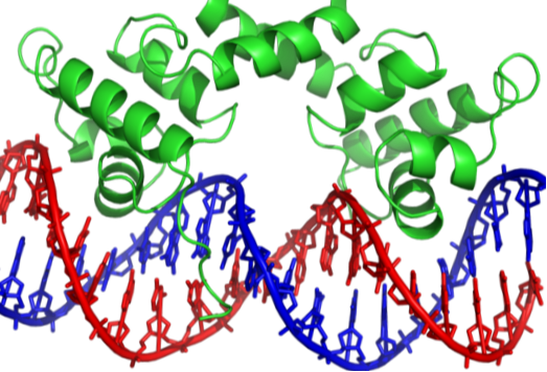
Molecular Function: This describes the molecular-level activity that a gene's protein or RNA products are associated with. Some examples of a molecular function of a gene might include regulation of gene expression, catalytic activity, or transporter activity. Essentially, this is the molecular role of the protein within the cell [1,2].
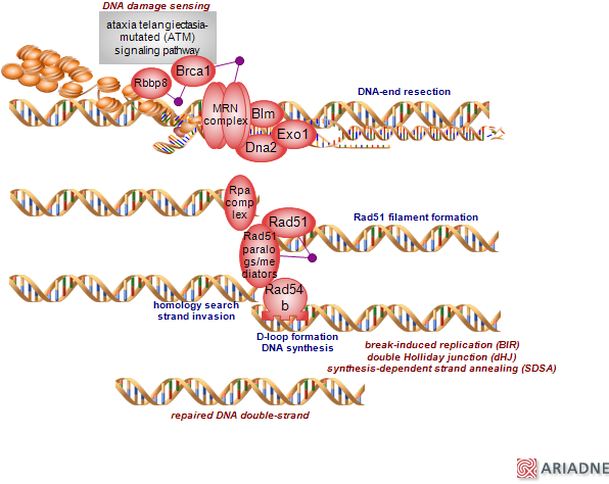
Biological Processes: Biological processes can be thought of as the next level up from molecular functions, as it describes a process that is powered by more than one molecular function within the cell. Some examples of biological processes terms include signal transduction, response to hypoxia, and DNA repair [1,2].
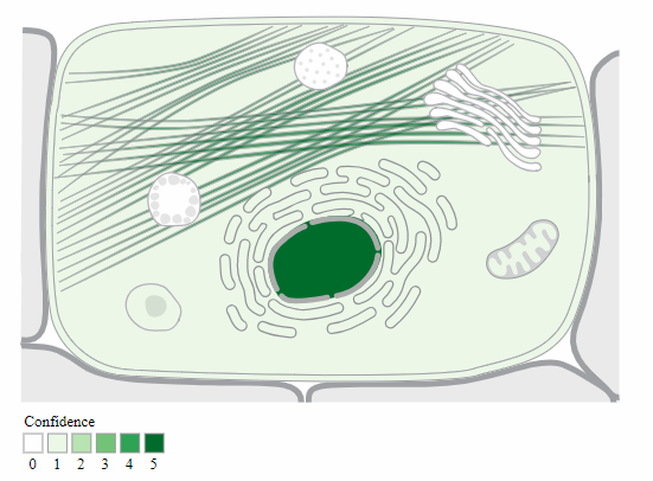
Cellular Component: While other aspects of gene ontology designate the function of a gene, the cellular component describes where the protein can be found within the anatomy of the cell. A cellular component term may reference large cellular structures such as the nucleus, the plasma membrane, or the mitochondria. However, cellular components can also refer to molecular structures, such as a specific protein complex (ie the proteasome) [1,2].
With numerous resources that can return the GO for a particular gene, researchers can quickly understand what role their gene plays within the cell.
Molecular Function: This describes the molecular-level activity that a gene's protein or RNA products are associated with. Some examples of a molecular function of a gene might include regulation of gene expression, catalytic activity, or transporter activity. Essentially, this is the molecular role of the protein within the cell [1,2].
Biological Processes: Biological processes can be thought of as the next level up from molecular functions, as it describes a process that is powered by more than one molecular function within the cell. Some examples of biological processes terms include signal transduction, response to hypoxia, and DNA repair [1,2].
Cellular Component: While other aspects of gene ontology designate the function of a gene, the cellular component describes where the protein can be found within the anatomy of the cell. A cellular component term may reference large cellular structures such as the nucleus, the plasma membrane, or the mitochondria. However, cellular components can also refer to molecular structures, such as a specific protein complex (ie the proteasome) [1,2].
With numerous resources that can return the GO for a particular gene, researchers can quickly understand what role their gene plays within the cell.
Cellular Component
|
Cellular Component annotations for RAD51D obtained from QuickGO:
|
Biological Processes
|
Biological Processes annotations for RAD51D obtained from QuickGO:
|
Molecular Function
|
Biological Processes annotations for RAD51D obtained from QuickGO:
|
Conclusions
The gene ontology of RAD51D supports its essential role in the homologous recombination process, both for DNA repair, as well as the generation of genetic diversity that occurs in meiosis. Likewise, there are some important aspects of the RAD51D that are essential for its ability to bind to damaged DNA. However, it is interesting to note in the absence of DNA damage, the RAD51D protein co-localizes with centrosomal elements outside the nucleus, suggesting the possibility of a regulatory mechanism to induce RAD51D nuclear location in the presence of DNA damage [3].
References:
1. An Introduction to the Gene Ontology. Retrieved from ftp://ftp.geneontology.org/go/www/GO.doc.shtml
2. GO Consortium. Gene Ontology overview. Retrieved from http://geneontology.org/docs/ontology-documentation/
3. Cappelli, E., Townsend, S., Griffin, C., Thacker, J. (2011). Homologous recombination proteins are associated with centrosomes and are required fro mitotic stability. Experimental Cell Research, 317(8), 1203-1213. Retrieved from https://www-sciencedirect-com.ezproxy.library.wisc.edu/science/article/pii/S0014482711000346
1. An Introduction to the Gene Ontology. Retrieved from ftp://ftp.geneontology.org/go/www/GO.doc.shtml
2. GO Consortium. Gene Ontology overview. Retrieved from http://geneontology.org/docs/ontology-documentation/
3. Cappelli, E., Townsend, S., Griffin, C., Thacker, J. (2011). Homologous recombination proteins are associated with centrosomes and are required fro mitotic stability. Experimental Cell Research, 317(8), 1203-1213. Retrieved from https://www-sciencedirect-com.ezproxy.library.wisc.edu/science/article/pii/S0014482711000346
References - Images:
Header: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5793732/