This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Introduction
Protein interactions, or how proteins physically associate with one another, can be informative for their role in cellular processes. Thanks to advances in the field of proteomics (the study of all the proteins present in a given system), researchers have identified an unfathomable quantity of protein-protein interactions. However, one can imagine the difficulty in sifting through this data publication by publication during research. Instead, multiple organizations have constructed databases that are curated with protein interacion data. Through this publicly available data, scientists have developed algorithms to construct Protein Interaction Networks which can mathematically quantify the physical contact between two proteins, or the strength of their interaction [1]. Furthermore, these algorithms possess predictive power, allowing users to uncover probable physical relationships that have not yet been experimentally verified [1].
Protein Interaction Networks are invaluable to researchers for a number of reasons. For example, there are countless proteins whose biological function have yet to be elucidated. Since these physical protein interactions serve a functional purpose in biology, we can learn quite a bit about what a protein does given the roles of its interactors. This "guilt by association" approach allows researchers to assign putative biological roles to proteins of interest that can be experimentally verified, leading to a small, but important contribution to our knowledge of the proteome [1]. Furthermore, using protein interactions allows for researchers to dissect signaling pathways, and identify proteins that come together to form large complexes that serve important functions within the cell [1].
Protein Interaction Networks are invaluable to researchers for a number of reasons. For example, there are countless proteins whose biological function have yet to be elucidated. Since these physical protein interactions serve a functional purpose in biology, we can learn quite a bit about what a protein does given the roles of its interactors. This "guilt by association" approach allows researchers to assign putative biological roles to proteins of interest that can be experimentally verified, leading to a small, but important contribution to our knowledge of the proteome [1]. Furthermore, using protein interactions allows for researchers to dissect signaling pathways, and identify proteins that come together to form large complexes that serve important functions within the cell [1].
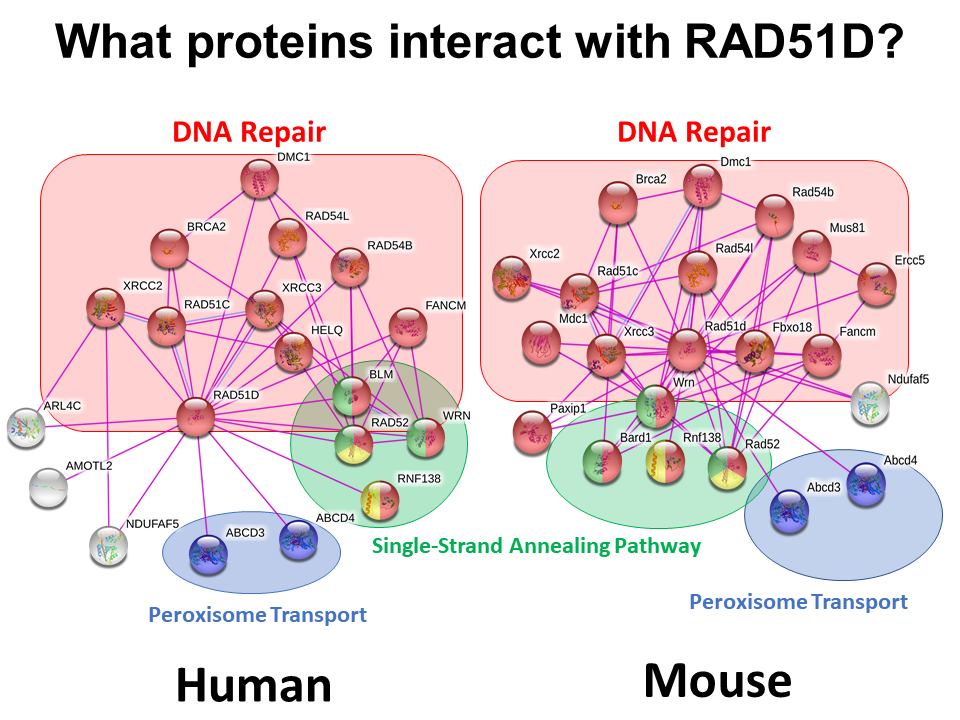
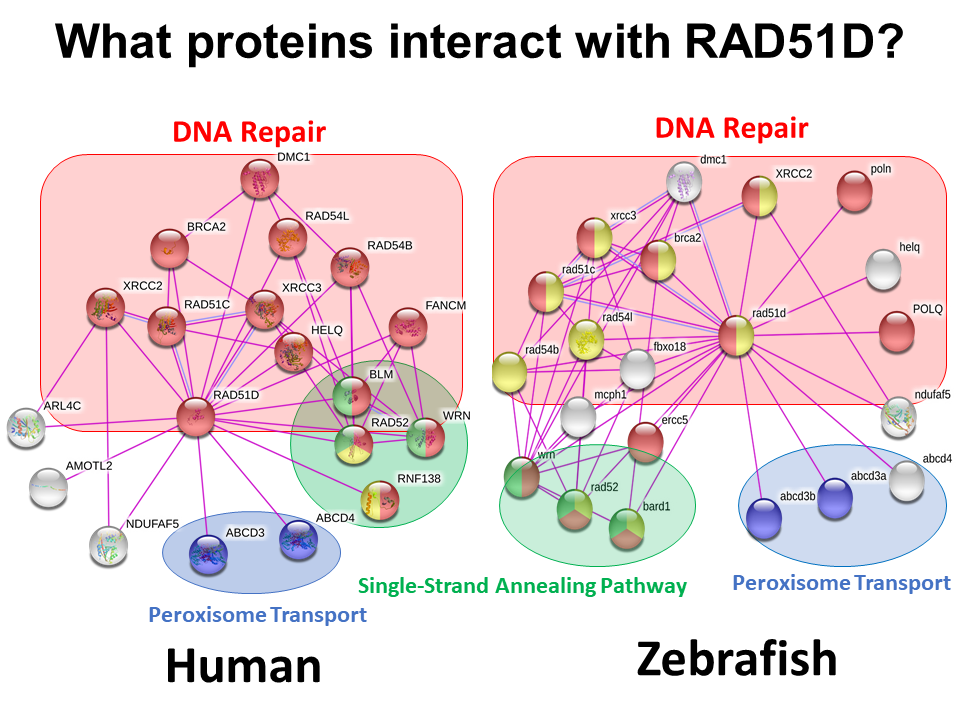
Figure 1: Protein interaction networks for RAD51D in humans, as well as the model organisms mice and zebrafish generated using the STRING Database. . The purple lines demonstrate protein interactions that have been reported in the literature.
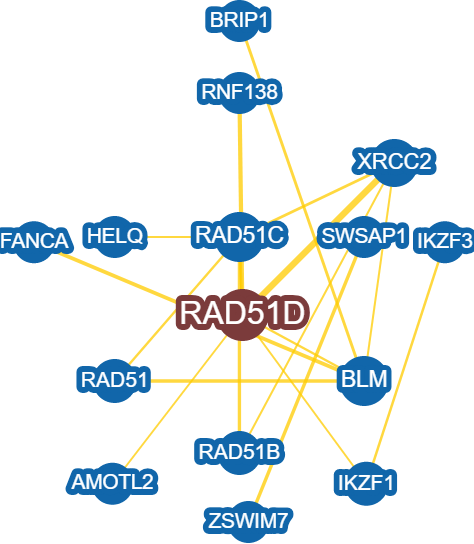
Figure 2: Predicted protein interaction network of RAD51D, generated using BioGRID.
Conclusions
Upon examination of the RAD51D protein networks in humans and the model organisms mice and zebrafish, there were a number of conserved interactions between the species. Specifically, RAD51D interacts with the BCDX2 complex components RAD51C and XRCC2 in each of the organisms of interest, emphasizing the importance of its function in maintaining genome integrity and diversity. However, there were some unexpected associations in both the human and mouse protein networks. For one, three proteins involved in regulating metabolism were identified, two of which function in transporting vitamin B12 or lipids into the peroxisome organelle. Furthermore, NDUFAF5 is an important regulator of complex 1, a protein that is essential for the generation of ATP through the electron transport chain of the mitochondria. Given that metabolic pathways in both of these organelles are affected in cancer, it will be interesting to see if RAD51D deficiencies predispose individuals to cancer independently of DNA repair.
Additionally, an interesting result was identified in the protein interaction of the zebrafish ortholog of RAD51D, where RAD51D was shown to interact with three proteins that regulate the single-stranded DNA repair pathway. Given that two of these three identified SSA regulators are present in both zebrafish and mice, it is likely that RAD51D directly interacts with RAD52, and BARD1 to regulate homologous repair in mammalian cells.
Additionally, an interesting result was identified in the protein interaction of the zebrafish ortholog of RAD51D, where RAD51D was shown to interact with three proteins that regulate the single-stranded DNA repair pathway. Given that two of these three identified SSA regulators are present in both zebrafish and mice, it is likely that RAD51D directly interacts with RAD52, and BARD1 to regulate homologous repair in mammalian cells.
References
1. EMBL-EBI. Protein-Protein Interaction Networks. Retrieved from https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
References - Images
Figure 1: https://string-db.org/
Figure 2: https://thebiogrid.org/
Header: http://www.mpip-mainz.mpg.de/40475/Nanoparticle_Protein_Interactions
Figure 2: https://thebiogrid.org/
Header: http://www.mpip-mainz.mpg.de/40475/Nanoparticle_Protein_Interactions