This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Introduction
What is Homology? [1,2]
Thanks to the recent advances in DNA sequencing, the genomes of over 42,746 species are publicly available through NCBI. With these resources, biologists everywhere are able to compare the biological features of thousands of species. When comparing genomes, biologists often search for homology, where closely related genes, proteins, or structures originate from a shared evolutionary origin, known as a common ancestor [1]. Take for example the forearms of the species depicted at the top of this page. While the different bone structures are conserved across the different species, the function of the individual bones can be variable; however, they all diverged from a single common ancestor [2].
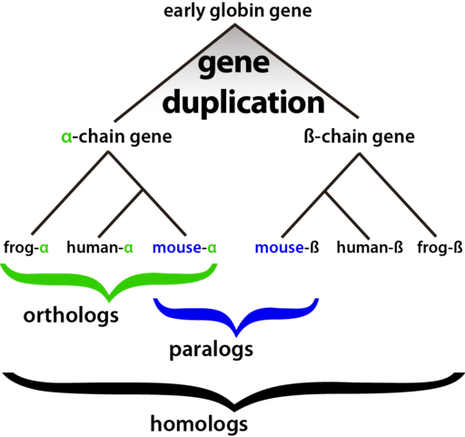
Orthologs vs Paralogs [1,2,3]Furthermore, homologs can be broken down into subcategories that better explain a gene or proteins evolutionary history. Shown to the right is an example of the horology of the globin gene, which is essential for the transport of oxygen throughout tissues. Homologs that occupy the same genetic loci, and have diverged from the same common ancestor are known as orthologs. For example, we can see in the figure to the right that frogs, humans, and mice share a alpha-chain gene that descended from the same early globin gene, making them orthologous to one-another. In contrast, we can also identify genes that originate from the expansion of a given gene. Homologs that result from gene duplication are known as paralogs. Returning to the globin gene example, the early globin gene experienced a gene duplication event, creating the alpha-chain and beta-chain globin genes. Since the mice alpha and beta globin genes originated from an ancestral duplication event they are classified as paralogs.
|
How Are Homologs Typically Identified?
Interactive explanation of BLAST to be added in the future!
RAD51D Homologs
|
Homo sapiens
(Human) |
Mus musculus
(Mouse) |
|
Pan troglodytes
(Common Chimpanzee) |
Macaca mulatta
(Rhesus Monkey) |
Rattus norvegicus
(Brown Rat) |
|
RAD51D (Isoform X1)
Accession: XP_001111649.2 Length: 431 Amino Acids 97% Identity Canis lupus familiaris (Dog) |
|
RAD51D (Isoform X1)
Accession:NP_001039769.1 Length: 326 Amino Acids 82% Identity Danio rerio (Zebrafish) |
RAD51D (Isoform X1)
Accession: XP_548263.2 Length: 328 Amino Acids 85% Identity Xenopus tropicalis (Western Clawed Frog) |
RAD51D
Accession: NP_001185575.1 Length 327 Amino Acids 67% Identity Drosophila melanogaster (Fruit Fly) |
|
rad51d
Accession: NP_996959.1 Length: 327 Amino Acids 55% Identity Arabidopsis thaliana (Thale Cress) |
rad51d
Accession: NP_001005687.1 Length: 320 Amino Acids 57% Identity Medicago truncatula (Barrel Medic) |
|
Escheria coli
(E. coli) |
Saccharomyces cerevisiae
(Baker's Yeast) |
Caenorhabditis elegans
(Roundworm) |
Conclusions
RAD51D is a particularly interesting gene to study due to its complicated evolutionary history. As a member of the RAD51 family, it is believed to have originated from a gene duplication of an ancestral RAD51-like protein. Thus, making RAD51D a paralog to the conserved RAD51 protein. Likewise, RAD51D also has orthologs that are conserved across multiple kingdoms, including both plant and animal species. Moving forward, the protein structure of the RAD51D protein homologs will be analyzed using phylogenetic and protein domain analyses in order to determine what structures are unique to organisms that have developed vertebrate ovaries.
References
1. Griffiths, A. J. F., Wessler, S. R., Carroll, S. B., Doebley, J. (2015). Introduction to Genetic Analysis (11th ed., pp. 528-529). New York, NY: W.H. Freeman and Company.
2. Time Scavengers Blog. Homology. Retrieved from https://timescavengers.blog/evolution/homology/
3. Bitesize Bio. Homology Terminology: Never Say the Wrong Word Again https://bitesizebio.com/26762/homology-terminology-never-say-wrong-word/
References - Images
Background: Homologous Structures
| FASTAS | |
| File Size: | 6 kb |
| File Type: | txt |