This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Introduction
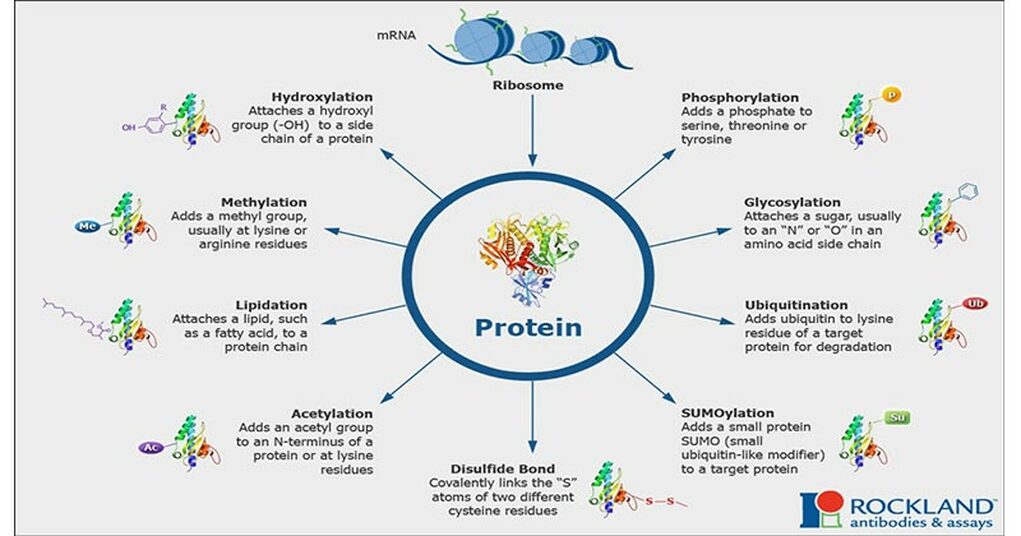
A proteins activity in a cell cannot be determined by its expression alone. Often, proteins are regulated by Post-Translational Modifications. These modifications can be chemical in nature, such the addition of a phosphoryl or ubiquitin group to specific residues found on the polypeptide. Additionally, post-translational modifications can involve cleavage of the protein at different sites, leading to peptides of a different size and structure. These protein modifications can be used for a variety of purposes in the cell. For example, phosphorylation often exerts regulatory effects on a protein, causing conformational changes that alter its activity on a particular substrate. Likewise, ubiquitination of specific amino acid residues often signals for the degradation of select proteins.. Finally, post-translational modifications can be important for helping a protein to fold correctly within the cell, or localize its target to a particular location within the cell. Clearly, post-translational modifications can exert an important impact on how a protein functions within the cell.
Figure 1: A depiction of various protein post-translational modifications.
Results
Figure 2: Predicted phosphorylation sites of the RAD51D protein in humans (A), mice (B), and zebrafish (C), generated using NetPhos. Only phosphorylation site with scores with high-confidence (>0.8) are displayed.
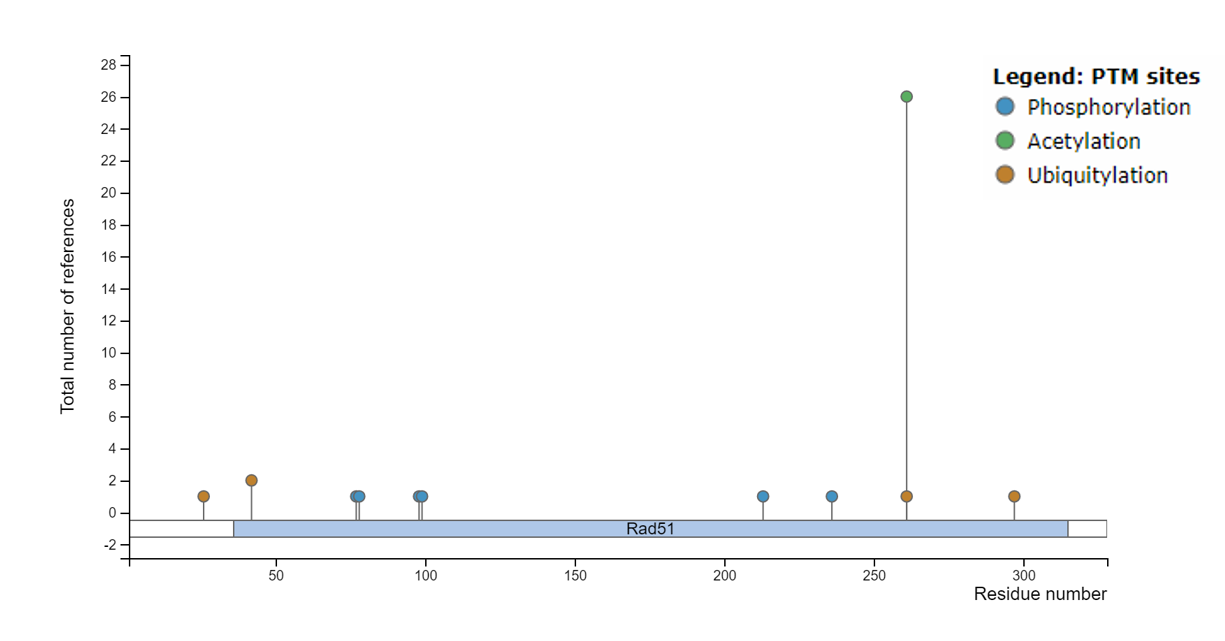
Figure 3: Predicted post-translational modifications sites for the RAD51D protein predicted by PhosphoSitePlus.
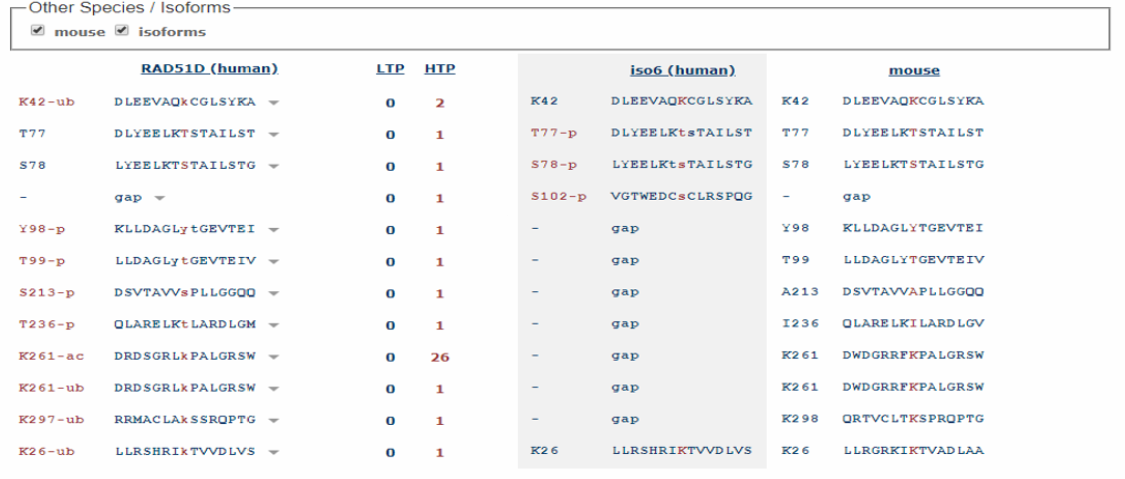
Figure 4: Post-translational modification sites on the RAD51D protein shared between human and mice. Generated using PhosphoSitePlus
Conclusions
After examining the post-translational modifications across humans, mice, and zebrafish, it was interesting to note the likelihood of a phosphorylation site on serine 46 in each of the RAD51D orthologs. Furthermore, PhosphoSitePlus identified the presence of several ubiquitin sites, as well as acetylation. Given the fact that both of these modifications can regulate the activity of a given protein, it will be interesting to examine if mutations in residues K261, K42, or S46 affect DNA repair activity. Furthermore, the high number of references of acetylation at K261 definitively establishes its role as an important post-translational modification for the RAD51D protein.
References - Images
Figure 1: https://rockland-inc.com/uploadedImages/ProductsStatic/post-translational-modifications-ptm.gif
Figure 2: http://www.cbs.dtu.dk/services/NetPhos/
Figure 3: https://www.phosphosite.org/homeAction.action
Figure 4: https://www.phosphosite.org/homeAction.action
Header: https://www.biocompare.com/Editorial-Articles/347122-On-the-Hunt-for-Post-Translational-Modifications/
Figure 2: http://www.cbs.dtu.dk/services/NetPhos/
Figure 3: https://www.phosphosite.org/homeAction.action
Figure 4: https://www.phosphosite.org/homeAction.action
Header: https://www.biocompare.com/Editorial-Articles/347122-On-the-Hunt-for-Post-Translational-Modifications/