This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Introduction
What are Protein Domains? [1]
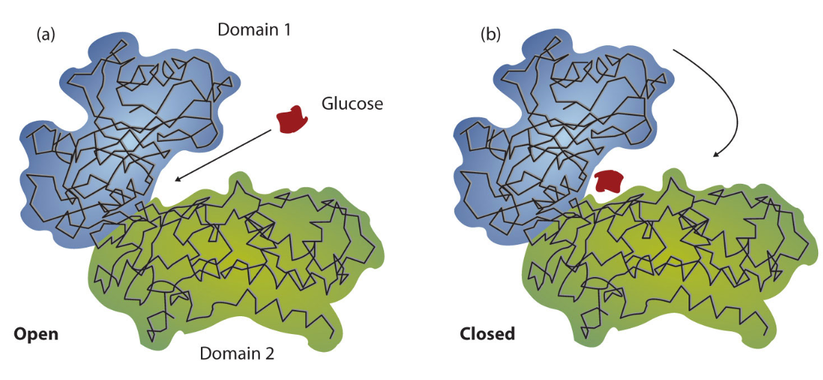
The three-dimensional structures of proteins are an essential determinant of the reactions and chemical processes they catalyze within a cell. Acting as distinct structural components of a protein, domains carry out specific and independent functions [1]. The function of a protein domain can vary based on its structure; however, some examples include the binding of small molecules, physical interactions with other proteins, or catalyzing a specific chemical reaction. Likewise, the amount of domains contained within a protein is variable; some proteins contain multiple domains, whereas smaller proteins exist as a single domain [1]. Interestingly, these domains often retain their shape even when chemically separated from the rest of the protein structure, emphasizing their ability to act independently [1]. An example of a protein with multiple domains is shown below.
Shown above is a diagram of the two domains of the enzyme Hexokinase. Notice how each of the two domains exist as a distinctive unit of the protein's structure.
Results
Protein Domains Present in RAD51D
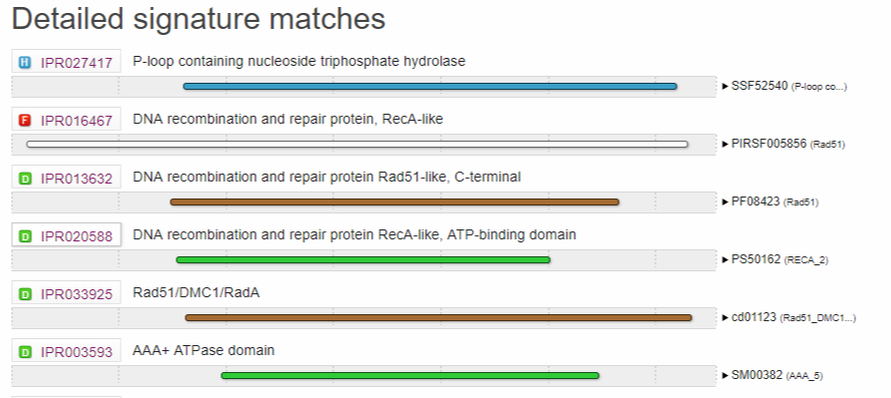
Figure 1A: Domains present in the Human RAD51D protein. Results were obtained from Interpro, using isoform 1 (canonical transcript) as the input query.
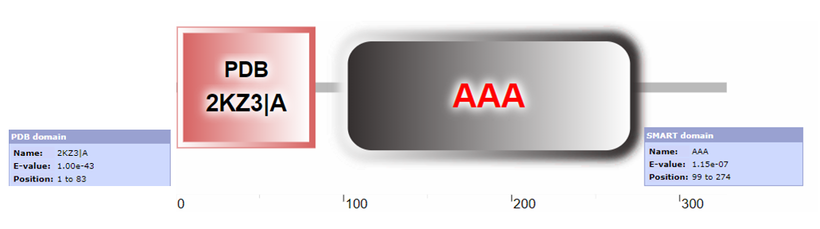
Figure 1B: Domains present in the Human RAD51D protein. Results were obtained from SMART, using isoform 1 as the input query. Due to overlap with the AAA domain, SMART is unable to display the RAD51 domain, but was a significant result returned by the SMART query.
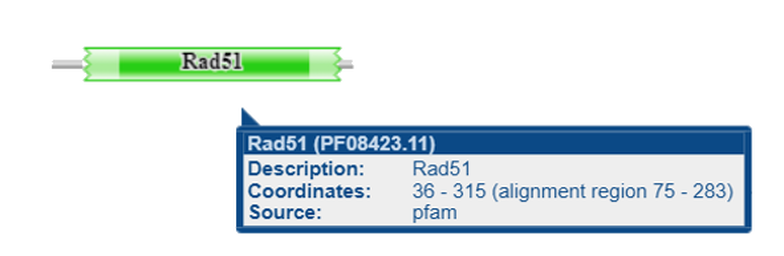
Figure 1C: Domains present in the Human RAD51D protein. Results were obtained from Pfam, using isoform 1 as the input query (evalue cutoff = 1e-5)
|
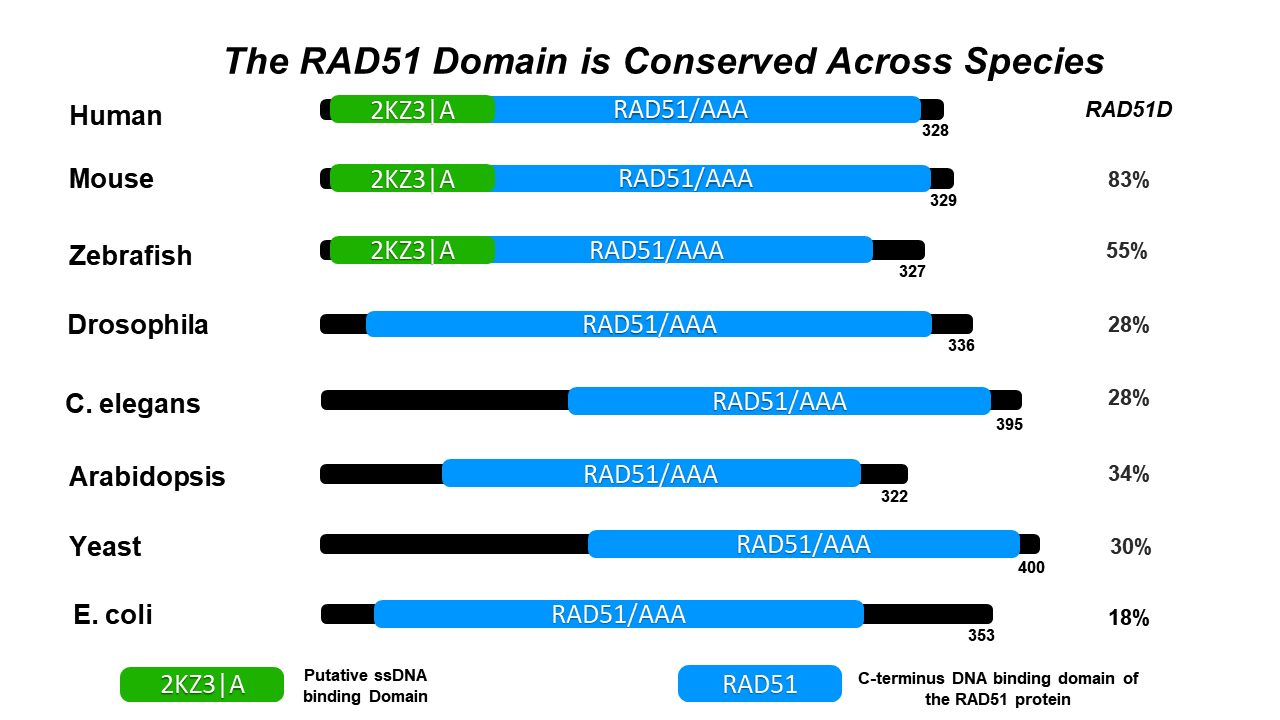
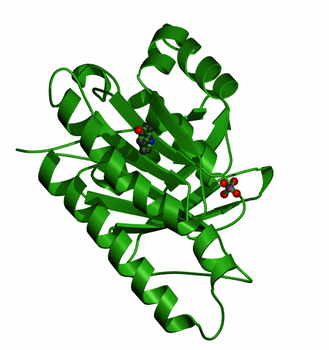
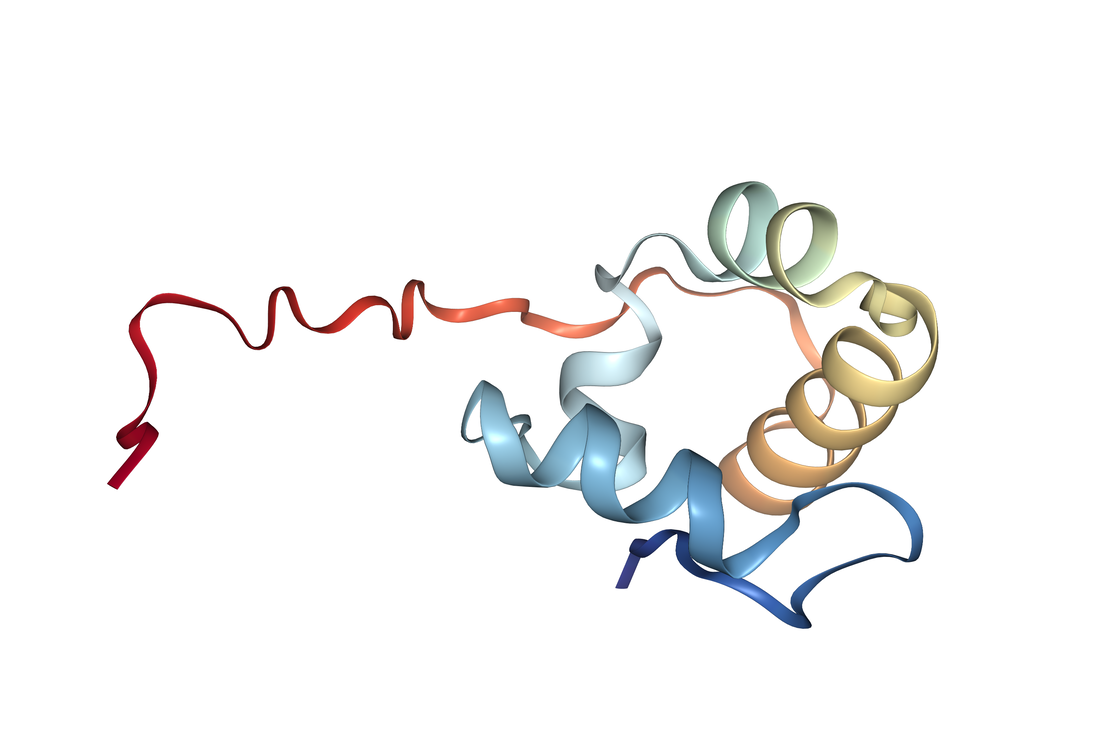
RAD51 Family Domain [2]: Shown to the right is the crystal structure of the DNA recombination and repair protein RAD51. From single-celled yeast, to humans, the RAD51 protein is highly conserved across multiple eukaryotic species. Both RAD51 and RAD51-like proteins comprise the RAD51 protein family, all of which share a domain near the c-terminus that is essential for DNA binding. Members of the RAD51 family also share a AAA domain (more information below) that is important for its DNA binding capabilities. Furthermore, the RAD51 family are homologous to the bacterial RecA, and the archael RadA, both of which also function in homologous repair and recombination [2]. |
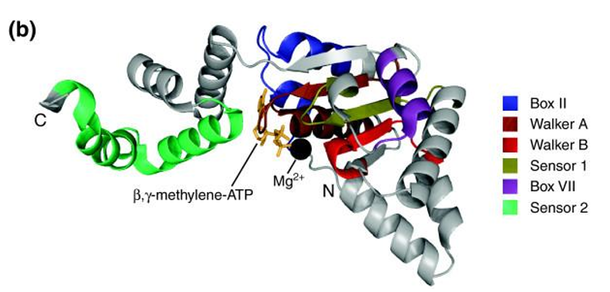
ATPases Associated With Diverse Cellular Activities (AAA) Domain [3]: AAA ATPases are a large protein family that function in a variety of essential biological processes, such as DNA-replication, cell-cycle regulation, protein assembly, and degradation. To accomplish these processes, AAA proteins bind and hydrolyze (break) the chemical bonds of the high-energy molecule ATP. By doing so, AAA proteins convert the energy released from ATP into mechanical energy that induces changes in their protein structure. Through these conformational changes, AAA protein domains can exert mechanical force on their macromolecule substrates. Furthermore, these AAA proteins share some important structural elements such as the Walker A/B motifs as well as ATP binding sites [3].
RAD51D Motifs/Conserved Residues
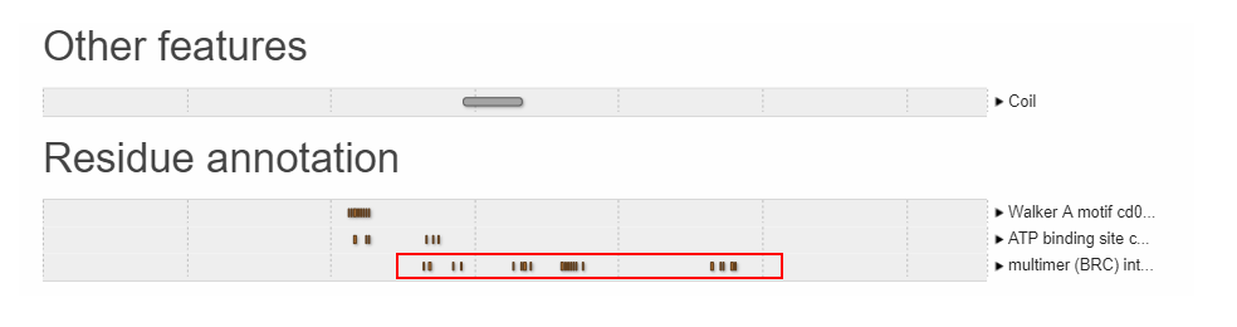
Figure 2: Annotated motifs and features of the RAD51D protein identified using interpro.
Conclusions
The RAD51/AAA domain is highly conserved across multiple model organisms that possess a RAD51D homolog. This emphasizes the essential roleAD51 family proteins have in the homologous recombination and repair pathways. Furthermore, a putative ssDNA binding protein has been identified in vertebrates, raising the possibility of an important domain for signaling DNA damage. Additionally, the N-terminal domain may play an essential role for hormonal regulation of ovarian cells and will be interesting for downstream analyses. Interestingly, like the BRCA2 protein, RAD51D also contains BRC interface repeats that are important for assembly of the RAD51 protein filament. In fact, it has been shown that mutations in the BRC domain of BRCA2 confered higher risk to ovarian cancer instead of breast cancer [5]. Using this information, I will compare the protein sequences of RAD51D in species with vertebrate ovaries to those of non-vertebrate origin, to determine if amino acids are conserved exclusively in vertebrate lineages.
References:
1. Nelson, D., Cox, M. (2017). Lehninger Principles of Biochemistry (7th Edition). New York, NY: W.H. Freeman and Company.
2. Pfam. Summary: RAD51. Retrieved from: https://pfam.xfam.org/family/Rad51#tabview=tab0
3. Pfam. Summary: ATPase family associated with various cellular activities (AAA). Retrieved from: https://pfam.xfam.org/family/AAA#tabview=tab0
4. Mee Kim, Y., Choi, B. S. (2010). Structural and functional characterization of the N-terminal domain of human Rad51D. The International Journal of Biochemistry & Cell Biology, 43(3), 416-422. doi: https://doi.org/10.1016/j.biocel.2010.11.014
5. Carreira, A., & Kowalczykowski, S. C. (2011). Two classes of BRC repeats in BRCA2 promote RAD51 nucleoprotein filament function by distinct mechanisms. Proceedings of the National Academy of Sciences of the United States of America, 108(26), 10448-53.
1. Nelson, D., Cox, M. (2017). Lehninger Principles of Biochemistry (7th Edition). New York, NY: W.H. Freeman and Company.
2. Pfam. Summary: RAD51. Retrieved from: https://pfam.xfam.org/family/Rad51#tabview=tab0
3. Pfam. Summary: ATPase family associated with various cellular activities (AAA). Retrieved from: https://pfam.xfam.org/family/AAA#tabview=tab0
4. Mee Kim, Y., Choi, B. S. (2010). Structural and functional characterization of the N-terminal domain of human Rad51D. The International Journal of Biochemistry & Cell Biology, 43(3), 416-422. doi: https://doi.org/10.1016/j.biocel.2010.11.014
5. Carreira, A., & Kowalczykowski, S. C. (2011). Two classes of BRC repeats in BRCA2 promote RAD51 nucleoprotein filament function by distinct mechanisms. Proceedings of the National Academy of Sciences of the United States of America, 108(26), 10448-53.
References - Images
Figure 1A: https://www.ebi.ac.uk/interpro/
Figure 1B: http://smart.embl-heidelberg.de/
Figure 1C: https://pfam.xfam.org/
Figure 2: https://www.ebi.ac.uk/interpro/
Header Background: 3D crystal structure of pyruvate kinase, generated using JSmol. Retrieved from http://proteopedia.org/wiki/index.php/Pyruvate_Kinase
Figure 1B: http://smart.embl-heidelberg.de/
Figure 1C: https://pfam.xfam.org/
Figure 2: https://www.ebi.ac.uk/interpro/
Header Background: 3D crystal structure of pyruvate kinase, generated using JSmol. Retrieved from http://proteopedia.org/wiki/index.php/Pyruvate_Kinase